1Department of Oral Microbiology, College of Dentistry, Kyung Hee University
2Department of Dentistry, Graduate School of Kyung Hee University
Correspondence to Eun-Young Jang, Department of Oral Microbiology, College of Dentistry, Kyung Hee University, 26 Kyungheedae-ro, Dongdaemun-gu, Seoul-si, 02447, Korea. Tel: +82-2-961-0598, E-mail: ur1010@khu.ac.kr
Volume 25, Number 3, Pages 189-98, June 2025.
J Korean Soc Dent Hyg 2025;25(2):189-98. https://doi.org/10.13065/jksdh.2025.25.3.1
Received on April 09, 2025, Revised on May 09, 2025, Accepted on May 13, 2025, Published on June 30, 2025.
Copyright © 2025 Journal of Korean Society of Dental Hygiene.
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License(http://creativecommons.org/licenses/by-nc/4.0).
Objectives: This study aimed to evaluate the influence of eluents from direct-printed aligners (DPAs) on Streptococcus mutans (S. mutans), a major cariogenic bacterium, by examining bacterial growth, biofilm formation, and transcriptomic changes. Methods: Eluents were collected by immersing 3D-printed aligners in artificial saliva under simulated oral conditions. These eluents were then used to treat S. mutans, and bacterial proliferation was measured by optical density at 600 nm (OD600 ). Biofilm formation was quantified using crystal violet staining, and ultrastructural alterations were assessed via scanning electron microscopy (SEM). Transcriptomic shifts were identified by RNA sequencing (RNA-seq). Results: DPA eluents reduced S. mutans growth and biofilm formation by 29-39% compared to controls (p<0.05). SEM revealed decreased biofilm without morphological alterations. RNAseq analysis detected 10 differentially expressed genes (DEGs), showing downregulated expression of quorum sensing (QS) and biofilm-associated genes, including lantibiotic-related and acyl carrier protein genes. Conversely, genes related to oxidative stress responses-such as aldo-keto reductases, short-chain dehydrogenases, and ATP-binding cassette transporters-were upregulated, suggesting activation of a stress-adaptive mechanism. Conclusions: While DPA eluents effectively suppress biofilm formation, the accompanying activation of oxidative stress responses may enhance bacterial resilience, raising concerns about potential long-term microbiome dysbiosis. Comprehensive microbial evaluations of orthodontic materials remain essential to ensure clinical safety and efficacy.
Biofilm, Direct-printed aligner, RNA sequencing, Streptococcus mutans, Transcriptome
The advancement of three-dimensional (3D) printing technology and biomaterials, computer-aided design (CAD) and manufacturing (CAM) have significantly influenced the field of orthodontics, particularly in the production of clear aligners and retainers [1]. Clear aligners offer several advantages over traditional fixed orthodontic appliances, including enhanced aesthetics, ease of use, and improved oral hygiene management [2,3]. Additionally, direct-printed aligners (DPAs) have garnered significant attention due to their environmental advantages and the customization potential of in-house fabrication systems [4,5].
However, their widespread adoption has been accompanied by an increasing number of reports of adverse clinical effects. Allareddy et al. [6] analyzed data from the Manufacturer and User Facility Device Experience database and identified 169 adverse events associated with Invisalign over a 10-year period from 2006. Reported cases ranged from mild reactions, such as skin rashes, to severe or potentially life-threatening complications, including breathing difficulties, sore throat, swelling of the throat, tongue, or lips, hives, anaphylaxis, and sensations of airway obstruction or laryngospasm. Willi et al. [7] detected urethane dimethacrylate in all tested DPA samples, with concentrations ranging from 29 to 96 μg/L after immersion in distilled water (DW) for one week. When considered together, these studies point to potential concerns regarding the safety of DPAs use. Given that patients are typically instructed to replace DPAs every 7–10 days, the repeated introduction of new aligners into the oral cavity may contribute to continuous exposure to residual chemicals, raising potential health concerns. If aligners are not systematically tested for material leaching or contain harmful chemical components, extended exposure could pose significant health risks, akin to repeated contact with a toxic substance.
Beyond potential cytotoxic effects, the accumulation of biofilm on DPAs can contribute to dental caries, periodontal disease, and other oral health complications [2,3,8]. Despite being easily removable, DPAs may promote biofilm formation by disrupting saliva flow and natural self-cleaning mechanisms when worn for extended periods [1,9,10]. Karkhanechi et al. [11] reported shifts in the periodontal microbiota following clear aligner usage, and more recently Yan et al. [9] has confirmed that biofilm accumulation on clear aligners is associated with microbiome alterations linked to dental caries. Additionally, the use of clear aligners has been shown to contribute to the colonization of Streptococcus mutans (S. mutans), further increasing the risk of caries in orthodontic patients [2].
Despite the growing interest in DPAs, studies investigating their interactions with S. mutans, a key contributor to dental caries development, remain limited. To address this gap, the present study aimed to evaluate the effects of DPAs on the growth and biofilm formation of S. mutans using eluent from DPAs. Additionally, we utilized RNA sequencing (RNA-seq) to analyze the associated gene expression profiles of S. mutans in response to DPAs.
The digital scan of the maxillary arch after non-extraction treatment was imported into the Direct Aligner Designer (DAD) software (Graphy, Seoul, South Korea) for the virtual aligner design. This design was then exported to the Uniz Dental software for 3D printing using the Uniz NBEE printer (Uniz, CA, USA) with Tera Harz TA-28 DAC resin (Graphy, Seoul, South Korea). Samples were prepared according to a protocol similar to the manufacturer’s instructions for aligner fabrication. After printing with the Uniz NBEE printer (Uniz, CA, USA), the DPAs were removed from the platform and centrifuged for 6 minutes after removing the supports to eliminate residual monomers using a Tera Harz Spinner (Graphy, Seoul, South Korea). The DPAs were post-cured for 20 minutes using high-intensity LEDs and nitrogen generators to ensure curing in an oxygen-free environment using Tera Harz Cure (Graphy, Seoul, South Korea).
Each DPA was cut into 28 pieces to minimize the amount of artificial saliva (Sigma-Aldrich, Cat. SAE0149) required for complete immersion. All pieces of the three DPAs were immersed in a 50 mL tube containing 24 mL of artificial saliva and continuously shaken in a shaking incubator set at 37°C and 130 rpm for 7 days to facilitate elution. Evaporative losses during the incubation period were compensated by the addition of DW. All eluent samples were sealed and stored at 4°C until further use in the experiment. The following types of eluents were prepared. E1DPA (Eluate from one DPA): After 7 days of incubation with the pieces from one DPA, the eluent was collected. One of the four E1DPA 50 mL tubes was stored, while the remaining three tubes were used to prepare E2DPA (Eluate from 2 DPAs). The second set of DPA pieces was added to the eluent collected from E1DPA and incubated for another 7 days. After incubation, one tube was stored as E2DPA, while the remaining two tubes were used to prepare E3DPA (Eluate from 3 DPAs). The third set of DPA pieces was added to the eluent collected from E2DPA and incubated for 7 days. After incubation, one tube was stored, while the remaining tube was used to prepare E4DPA (Eluate from 4 DPAs): The fourth set of DPA pieces was added to the eluent collected from E3DPA and incubated for 7 days. The final eluent was collected and stored as E4DPA <Fig. 1>.
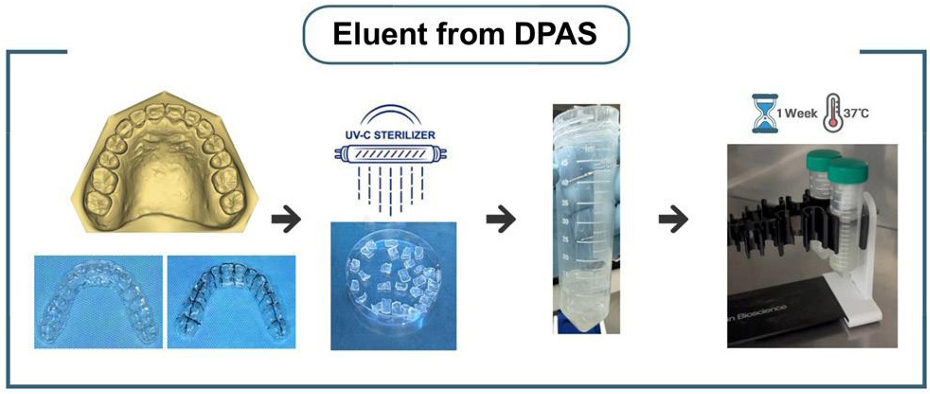
Fig. 1. Preparation of eluents from DPAs
DPAs were cut into smaller pieces to facilitate elution and sterilized with UVC. The DPA pieces were then placed in tubes containing artificial saliva and incubated at 37°C with continuous shaking for one week. The final eluates were stored at 4°C.
S. mutans GS-5 was used as the experimental strain. The bacterium was inoculated into Brain Heart Infusion (BHI; Becton, Dickinson and Company, Sparks, MD, USA) broth or BHI agar supplemented with Micro agar (Duchefa Biochemie, RV Haarlem, Netherlands). Cultures were incubated at 37°C under anaerobic conditions (5% CO2 , 5% H2 , 90% N2 ) using an anaerobic chamber (Coy Laboratory Products, Michigan, USA).
Biofilms were prepared with or without exposure to aligner eluents, as described above. The biofilms were then gently rinsed three times with physiological saline to remove non-adherent cells. Primary fixation was performed using 2.5% (wt/vol) glutaraldehyde prepared in 0.1 M phosphate buffer (pH 7.4), sterilized through a 0.22 µm filter, and incubated for 1 hour at room temperature. Secondary fixation was carried out using 1% (wt/vol) osmium tetroxide dissolved in 0.1 M phosphate buffer (pH 7.4) for an additional hour. After fixation, the biofilm was sequentially dehydrated using a graded ethanol series consisting of 25%, 50%, and 75% ethanol (one wash per concentration), followed by two washes with 100% ethanol to achieve complete dehydration. The dehydrated sample was dried using a freeze dryer (Ilshin Biobase Co. Ltd., Gyeonggi, South Korea) and coated with a thin layer of gold using a sputter coater (IB-3, Eiko, Tokyo, Japan) to enhance conductivity. The prepared sample was subsequently examined using a scanning electron microscope (model S-4700, Hitachi High Technologies America Inc., Pleasanton, CA, USA) operating at 10 kV.
Each of the control and experimental groups consisted of two biological replicates. The control group was treated with artificial saliva, while the experimental group was exposed to DPA eluents (E4DPA) for five hours. Total RNA was extracted using TRIzol™ Reagent (Invitrogen™, Cat. 15596026) and the RNeasy® Mini Kit (QIAGEN, Cat. 74104), according to the manufacturers’ instructions. Only high-quality RNA samples with an RNA Integrity Number (RIN) ≥7.0 were selected for RNA library construction. RNA libraries were constructed using the Illumina TruSeq Stranded mRNA Sample Prep Kit (Illumina, Inc., San Diego, CA, USA, #RS-122-2101). Bacterial rRNA was first removed from the samples using the NEBNext rRNA Depletion Kit (NEB, Cat. 850L). Subsequent to rRNA removal, the remaining RNA was subjected to cDNA synthesis and shearing as per the provided manufacturer’s instructions. The resulting product was then purified by PCR and concentrated to form the final cDNA library. Libraries were quantified using the KAPA Library Quantification Kit, designed for the Illumina sequencing platforms, and their quality assessed using the TapeStation D1000 ScreenTape (Agilent).
Paired-end reads were generated using the Illumina HiSeq Xten platform. Before analysis, Trimmomatic v0.38 was used to remove adapter sequences and low-quality bases from the raw data to generate high-quality reads. The trimmed reads were aligned to the S. mutans GS-5 genome obtained from the NCBI Genome Assembly database using the Bowtie program, and gene annotations were downloaded from the NCBI RefSeq database. The aligned data (BAM file format) were converted and indexed using SAMtools v1.17. After alignment, transcripts were assembled and quantified using the HTseq program. Gene-level expression profiles were quantified using the read counts from the aligned BAM files for each sample. Analyses were performed in R version 4.2.2. EdgeR’s exactTest identified significant genes based on raw counts, with genes selected for nonzero counts across samples. After RLE normalization to adjust for library size variations, differentially expressed genes (DEGs) were identified with a fold change ≥2 and a raw p<0.05.
The Shapiro-Wilk test was utilized to evaluate the normality of the data distributions in quantifying total bacterial growth and biofilm biomass. This test indicated non-normal distributions (p<0.05), prompting the use of the nonparametric Mann-Whitney U test for comparisons between groups. Results are depicted as medians and interquartile ranges (IQR). Box plots were generated to visually demonstrate the data distribution and to highlight differences between groups. Differences were considered statistically significant at p<0.05.
The effects of DPA eluents on the total growth of S. mutans were assessed by measuring the OD600 of the mixture of planktonic and biofilm cells. The quantification of S. mutans biofilm formation was evaluated using crystal violet staining. As shown in <Fig. 2A>, the total growth of S. mutans was significantly reduced when treated with the DPA eluents compared to the control (Ctrl) (p<0.0001). However, no statistically significant differences were observed among the aligner eluents (E1DPA, E2DPA, E3DPA, and E4DPA). Biofilm formation was visualized using crystal violet staining, showing that treatment with the aligner eluents decreased biofilm formation compared to the control <Fig. 2B>. Quantitative analysis showed a 29-39% reduction in S. mutans biofilm formation with the aligner eluents compared to the control <Fig. 2C>.
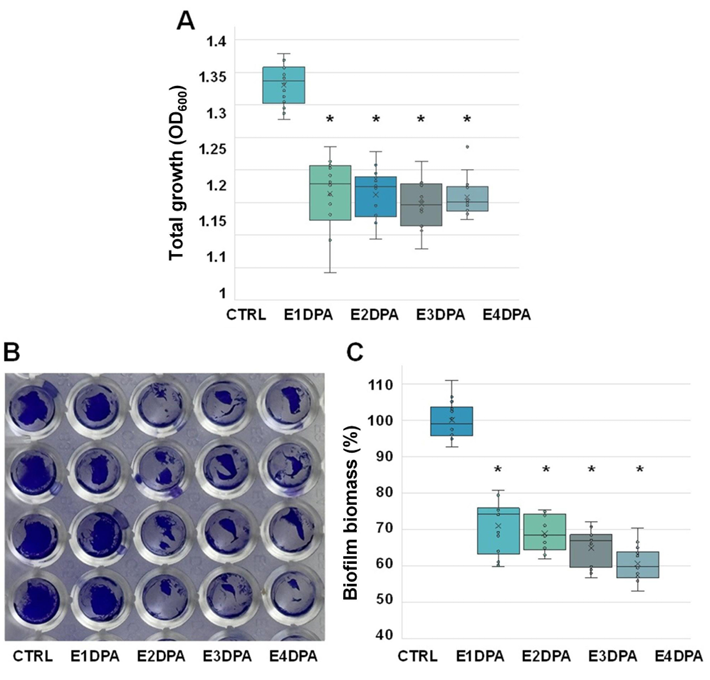
Fig. 2. Total growth and biofilm formation of S. mutans exposed to DPA eluents
(A) After 24 hours of incubation, the total growth of S. mutans was determined by measuring the OD600 . (B) Representative images of biofilm formation visualized by crystal violet staining after the aspiration of planktonic bacterial cells and spent medium. All experiments were conducted in triplicate (biological replicates) with a minimum of four technical replicates per condition. (C) Quantification of biofilm biomass based on crystal violet staining. After staining, the plate was washed three times with physiological saline and air-dried. Bound crystal violet was dissolved by the addition of 200 μL of 95% ethanol, followed by elution for 30 minutes. The OD600 of the eluate was subsequently measured. The box plot represents the biofilm biomass of S. mutans exposed to fresh artificial saliva (Ctrl) or DPA eluents. The graph illustrates the median, mean (denoted by ‘X’), interquartile range (IQR), and whiskers extending to 1.5 times the IQR, with outliers represented as individual points. Statistically significant differences were indicated by an asterisk for p<0.0001.
Exposure of S. mutans to fresh artificial saliva (Ctrl) or DPA eluents for 24 hours resulted in no observable changes in bacterial cell morphology. However, a notable reduction in biofilm cell density was observed. Remarkably, S. mutans exposed to E4DPA exhibited a pronounced decrease in the number of S. mutans biofilm cells after 24 hours of incubation <Fig. 3>.
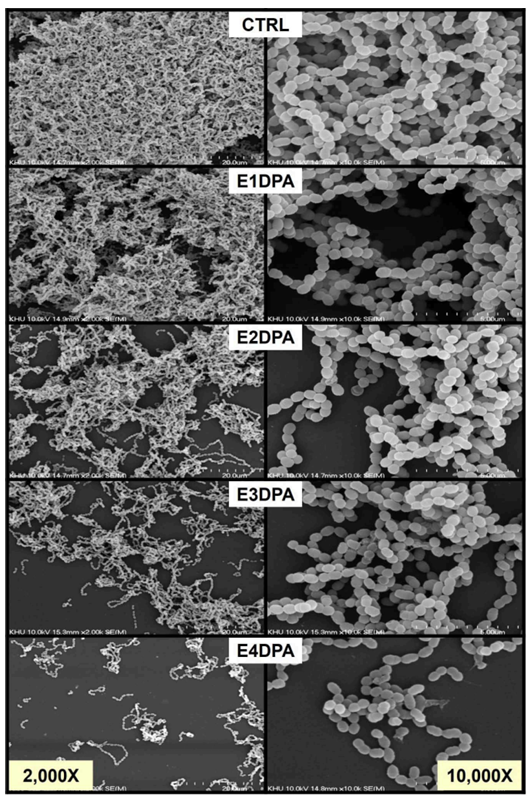

Fig. 3. Biofilm formation and ultrastructure of S. mutans unexposed or exposed to DPA eluents
S. mutans culture was adjusted to an OD600 of approximately 0.05 in BHI broth and then further incubated in 24-well plates containing fresh artificial saliva (Ctrl) and DPA eluents for 24 hours. SEM images of S. mutans biofilms were observed at magnifications of 2,000× (left row) and 10,000× (right row), operating at 10 kV.
Table 1. DEGs in S. mutans upon exposure to DPA eluents
테이블
Quantitative analysis of bacterial growth, measured by OD600 , revealed a significant decline in S. mutans growth in the presence of DPA eluents. SEM imaging further confirmed a marked attenuation in biofilm density, suggesting that DPA eluent exposure induces widespread cellular stress and disrupts regulatory pathways essential for biofilm formation and stability. Transcriptomic data further support these findings.
RNA-seq analysis identified 10 DEGs, with 4 upregulated and 6 downregulated following DPA eluent exposure. Among the downregulated genes were those encoding a lichenicidin alpha family lanthipeptide and acyl carrier protein, indicating suppression of pathways related to quorum sensing (QS), biofilm integrity, and membrane stability. Lantipeptides, a subclass of ribosomally synthesized and post-translationally modified peptides, include lantibiotics known for their antimicrobial properties. These compounds disrupt bacterial viability by forming membrane pores [12] and/or by inhibiting cell wall synthesis via binding to lipid II, a key intermediate in peptidoglycan biosynthesis [13,14]. In S. mutans, lantibiotics and other bacteriocin-like peptides are tightly regulated by the QS systems such as MutRS [15], which modulates microbial competition and biofilm homeostasis [16]. Bacteriocins governed by QS also contribute to biofilm structural maintenance by lysing competing bacteria and influencing the availability of extracellular DNA [17]. Moreover, QS systems coordinate both competence development and antimicrobial peptide production [18]. Therefore, the downregulation of lantibiotic-related genes may reflect disruptions in QS signaling, which could indirectly compromise the composition and resilience of the oral biofilm community.
Although direct evidence linking ACP to biofilm formation in S. mutans is limited, studies in Pseudomonas aeruginosa have shown that AcpP is essential for fatty acid biosynthesis and membrane lipid organization under oxidative stress [19]. Given the established role of the QS systems in coordinating biofilm development and cell-to-cell communication in both P. aeruginosa [20] and S. mutans [21], the observed downregulation of ACP following DPA eluent exposure may impair membrane stability by affecting QS signaling fidelity, thereby compromising biofilm structure.
In contrast, genes involved in oxidative stress responses were significantly upregulated. Notably, aldo-keto reductase and shortchain dehydrogenase/reductase family oxidoreductases-key enzymes in microbial detoxification and redox balance-were elevated [22,23]. These seemingly contradictory results may reflect the dual response of S. mutans to DPA eluent exposure, in which S. mutans activates stress-response enzymes as a compensatory response to mitigate oxidative damage caused by biofilm destabilization and altered lantibiotic regulation. Supporting this, the expression of ABC transporter genes also increased. ABC transporters facilitate cellular stress responses by exporting toxic metabolic byproducts and maintaining intracellular homeostasis [24]. Although ABC transporters are widely recognized for conferring antibiotic resistance by exporting drugs and toxic substances [25], their upregulation in this study likely reflects a generalized adaptation to DPA exposure, rather than a specific resistance mechanism.
Since this study focuses on transcriptome data, protein-level validation such as proteomic analysis will likely be required to confirm the functional relevance of the identified DEGs.
The interplay between the inhibition of biofilm formation, the regulation of QS, and oxidative stress adaptation has broader clinical implications. Mummolo et al. [26] proposed that orthodontic aligners, due to extended intraoral wear and creation of enclosed microenvironments, may promote microbial persistence by selecting for oxidative stress-resistant populations. Within the confined environment of aligners, such adaptations could exacerbate enamel demineralization and periodontal disease by enhancing bacterial tolerance to oxidative stress. Furthermore, suppression of QS-regulated lantipeptide production could shift the microbial balance, favoring colonization by non-S. mutans pathogens and altering disease progression. Previous study culturebased approach did not assess molecular adaptations such as reactive oxygen species (ROS)-detoxifying enzyme activity or stressresponsive gene expression. However, this study provides transcriptomic-level understanding into bacterial responses to DPA eluents. By revealing molecular changes associated with biofilm regulation, stress adaptation, and transporter activation, our findings address a critical gap in the mechanistic understanding of microbial-material interactions.
Taken together these findings highlight the complex and multifaceted microbial responses to orthodontic materials. As biofilm inhibition alone may not be sufficient to reduce pathogenicity if stress-adaptive mechanisms remain active, a more comprehensive strategy is warranted.
This study aimed to analyze the effects of DPA eluents on the growth, biofilm formation and transcriptomic changes of S. mutans, a major cariogenic bacterium.
1. DPA eluents significantly reduced biofilm formation in S. mutans and suppressed the expression of QS-related genes.
2. Nonetheless, the upregulation of oxidative stress-related genes suggests that S. mutans retains its ability to adapt to environmental stress and may continue to exhibit pathogenic characteristics.
3. Therefore, when evaluating orthodontic device materials, it is crucial to assess not only their physical and mechanical properties but also their impact on microbial gene expression.
In future studies, it will be essential to elucidate microbial adaptation mechanisms through integrated metagenomic, transcriptomic and proteomic analyses ultimately guiding the development of orthodontic materials that ensure both microbiological stability and clinical safety.
The author fully participated in the work performed and documented truthfully.
The authors declared no conflicts of interest.
None.
Non-human or animal research.
Data can be obtained from the corresponding author.
None.
1. Wu C, Mangal U, Seo JY, Kim H, Bai N, Cha JY, et al. Enhancing biofilm resistance and preserving optical translucency of 3D printed clear aligners through carboxybetaine-copolymer surface treatment. Dent Mater 2024;40(10):1575-83. https://doi.org/10.1016/ j.dental.2024.07.009
2. Rouzi M, Zhang X, Jiang Q, Long H, Lai W, LI X. Impact of clear aligners on oral health and oral microbiome during orthodontic treatment. Int Dent J 2023;73(5):603-11. https://doi.org/10.1016/j.identj.2023.03.012
3. Moradinezhad M, Montazeri EA, Ashtiani AH, Pourlotfi R, Rakhshan V. Biofilm formation of Streptococcus mutans, Streptococcus sanguinis, Staphylococcus epidermidis, Staphylococcus aureus, Lactobacillus casei, and Candida albicanson 5 thermoform and 3D printed orthodontic clear aligner and retainer materials at 3 time points: an in vitro study. BMC Oral Health 2024;24(1):1107. https://doi.org/10.1186 /s12903-024-04893-4
4. Eliades T, Zinelis S. Three-dimensional printing and in-house appliance fabrication: Between innovation and stepping into the unknown. Am J Orthod Dentofacial Orthop 2021;159(1):1-3. https://doi.org/10.1016/j.ajodo.2020.10.012
5. Alkhamees A. The new additive era of orthodontics: 3D-printed aligners and shape memory polymers-the latest trend-and their environmental implications. J Orthod Sci 2024;13(1):55. https://doi.org/10.4103/jos.jos_211_23
6. Allareddy V, Nalliah R, Lee MK, Rampa S, Allareddy V. Adverse clinical events reported during Invisalign treatment: analysis of the MAUDE database. Am J Orthod Dentofacial Orthop 2017;152(5):706-10. https://doi.org/10.1016/j.ajodo.2017.06.014
7. Willi A, Patcas R, Zervou SK, Panayi N, Schätzle M, Eliades G et al. Leaching from a 3D-printed aligner resin. Eur J Orthod 2023;45(3):244-9. https://doi.org/10.1093/ejo/cjac056
8. Tektas S, Thurnheer T, Eliades T, Attin T, Karygianni L. Initial bacterial adhesion and biofilm formation on aligner materials. Antibiotics 2020;9(12):908. https://doi.org/10.3390/antibiotics9120908
9. Yan D, Liu Y, Che X, Mi S, Jiao Y, Guo L, et al. Changes in the microbiome of the inner surface of clear aligners after different usage periods. Curr Microbiol 2021;78(5):566-75. https://doi.org/10.1007/s00284-020-02308-5
10. Low B, Lee W, Seneviratne CJ, Samaranayake LP, Hagg U. Ultrastructure and morphology of biofilms on thermoplastic orthodontic appliances in ‘fast’ and ‘slow’ plaque formers. Eur J Orthod 2011;33(5):577-83. https://doi.org/10.1093/ejo/cjq126
11. Karkhanechi M, Chow D, Sipkin J, Sherman D, Boylan RJ, Norman RG, et al. Periodontal status of adult patients treated with fixed buccal appliances and removable aligners over one year of active orthodontic therapy. Angle Orthod 2013;83(1):146-51. https:// doi.org/10.2319/031212-217.1
12. Chakraborty HJ, Gangopadhyay A, Datta A. Prediction and characterisation of lantibiotic structures with molecular modelling and molecular dynamics simulations. Sci Rep 2019;9(1):7169. https://doi.org/10.1038/s41598-019-42963-8
13. Hasper HE, Kramer NE, Smith JL, Hillman JD, Zachariah C, Kuipers OP, et al. An alternative bactericidal mechanism of action for lantibiotic peptides that target lipid II. Science 2006;313(5793):1636-7. https://doi.org/10.1126/science.1129818
14. Wiedemann I, Breukink E, van Kraaij C, Kuipers OP, Bierbaum G, de Kruijff B, et al. Specific binding of nisin to the peptidoglycan precursor lipid II combines pore formation and inhibition of cell wall biosynthesis for potent antibiotic activity. J Biol Chem 2001;276(3):1772-9. https:// doi.org/10.1074/jbc.M006770200
15. Wyllie RM, Jensen PA. The MutRS quorum-sensing system controls lantibiotic mutacin production in the human pathogen Streptococcus mutans. Proc Natl Acad Sci U S A 2025;122(7):e2421164122. https://doi.org/10.1073/pnas.2421164122
16. Shanker E, Federle MJ. Quorum sensing regulation of competence and bacteriocins in Streptococcus pneumoniae and mutans. Genes 2017;8(1):15. https://doi.org/10.3390/genes8010015
17. Cvitkovitch DG, Li YH, Ellen RP. Quorum sensing and biofilm formation in Streptococcal infections. J Clin Invest 2003;112(11):1626-32. https:// doi.org/10.1172/JCI20430
18. Reck M, Tomasch J, Wagner-Döbler I. The alternative sigma factor SigX controls bacteriocin synthesis and competence, the two quorum sensing regulated traits in Streptococcus mutans. PLoS Genet 2015;11(7):e1005353. https://doi.org/10.1371/journal.pgen.1005353
19. Ma JC, Wu YQ, Cao D, Zhang WB, Wang HH. Only Acyl Carrier Protein 1 (AcpP1) Functions in Pseudomonas aeruginosa Fatty Acid Synthesis. Front Microbiol 2017;8:2186. https://doi.org/10.3389/fmicb.2017.02186
20. Miranda SW, Asfahl KL, Dandekar AA, Greenberg EP. Pseudomonas aeruginosa Quorum Sensing. Adv Exp Med Biol 2022;1386:95-115. https:// doi.org/10.1007/978-3-031-08491-1_4
21. Senadheera D, Cvitkovitch DG. Quorum sensing and biofilm formation by Streptococcus mutans. Adv Exp Med Biol 2008;631:178-88. https:// doi.org/10.1007/978-0-387-78885-2_12
22. Elizabeth ME. Microbial aldo-keto reductases. FEMS Microbiol Lett 2002;216(2):123-31. https://doi.org/10.1111/j.1574-6968.2002.tb11425.x
23. Kavanagh KL, Jörnvall H, Persson B, Oppermann U. Medium- and short-chain dehydrogenase/reductase gene and protein families: The SDR superfamily: functional and structural diversity within a family of metabolic and regulatory enzymes. Cell Mol Life Sci 2008;65(24):3895-906. https://doi.org/10.1007/s00018-008-8588-y
24. Davidson AL, Dassa E, Orelle C, Chen J. Structure, function, and evolution of bacterial ATP-binding cassette systems. Microbiol Mol Biol Rev 2008;72(2):317-64. https://doi.org/10.1128/MMBR.00031-07
25. Lubelski J, Konings WN, Driessen AJ. Distribution and physiology of ABC-type transporters contributing to multidrug resistance in bacteria. Microbiol Mol Biol Rev 2007;71(3):463-76. https://doi.org/10.1128/MMBR.00001-07
26. Mummolo S, Nota A, Albani F, Marchetti E, Gatto R, Marzo G, et al. Salivary levels of Streptococcus mutans and Lactobacilli and other salivary indices in patients wearing clear aligners versus fixed orthodontic appliances: an observational study. PLoS One 2020;15(4):e0228798. https://doi.org/10.1371/journal.pone.0228798